Assembling the molecular machinery of coagulation: learning from the adaptive evolution of snake venom proteins
Hemostasis is regulated by the activity of macromolecular enzyme-cofactor complexes that require a negatively charged membrane surface for their assembly. Apart from spatially organizing the coagulation reactions upon vascular injury, the phospholipid surface-driven complexation also profoundly enhances the catalytic rate of protein substrate conversion. Suggestions as to how the membrane surface propagates the coagulation reactions include increased local reactant concentrations, induced conformational changes, or restricted protein movement. However, molecular details adequately describing this process are incomplete.
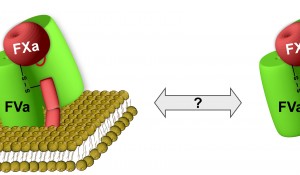
Here, we will provide new information on the molecular mechanisms that regulate the assembly of enzyme-cofactor complexes driving blood coagulation. In particular, we propose to study the role of the phospholipid surface in the complex formation of the blood coagulation cofactor factor Va and serine protease factor Xa that convert prothrombin to thrombin, a key step in blood coagulation. Variants of these proteins comprising improved procoagulant properties can be found in nature, with as most striking example the factor Va-Xa proteins expressed in the venom of the Australian common brown snake Pseudonaja textilis. We previously discovered that these proteins provide a sustained procoagulant stimulus via unique adaptations and seem to have evolved to circumvent the membrane-dependence in factors Va-Xa binding. By uncovering the molecular mechanism that allows these snake venom proteins to bind with high affinity in solution, we anticipate to be able to translate this to the human system, thereby gaining unprecedented insight.